Abstract
Background
Determination of estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2) status is standard for predicting prognosis and determining treatment options for patients with breast cancer. In 2010, the American Society of Clinical Oncology (ASCO) and College of American Pathologists (CAP) issued guidelines that tumors with ≥1 % positively staining cells should be considered ER positive. Here, we determined how this cutoff relates to molecular subtype.
Methods
Clinicopathological characteristics were compared between ER-negative, ER-positive, and low-ER-staining (1–10 %) tumors using chi-square analysis with P < 0.05 defining statistical significance. Gene expression data were generated for 26 low-ER-staining tumors, and their intrinsic subtype determined. Immunohistochemistry (IHC)-defined surrogate subtypes, using the threshold of positivity defined by ASCO/CAP guidelines, were compared with molecular subtypes.
Results
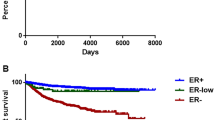
Low-ER-staining tumors were clinicopathologically more similar to ER-negative than to ER-positive tumors; 88 % of low-staining tumors were basal like or HER2 enriched. Only those tumors expressing 10 % ER-positive cells were classified as luminal A subtype.
Conclusions
Under ASCO/CAP guidelines, tumors with 1–10 % ER staining would be classified as ER positive, yet most are basal like or HER2 enriched and have pathological features similar to ER-negative tumors. Clinical trials seeking to treat tumors of ER-negative basal-like and/or HER2-enriched subtypes should thus not preclude enrollment based solely on results of ER immunohistochemistry. As ER status is a critical element in the choice of treatments for patients with breast cancer, it is imperative that the most effective method for classifying tumors be developed.


Similar content being viewed by others
References
Pietras RJ, Marquez-Garban DC. Membrane-associated estrogen receptor signaling pathways in human cancers. Clin Cancer Res. 2007;13:4672–6.
Aaltomaa S, Lipponen P, Eskelinen M, et al. Hormone receptors as prognostic factors in female breast cancer. Ann Med. 1991;23(6):643–8.
Dunnwald LK, Rossing MA, Li CI. Hormone receptor status, tumor characteristics, and prognosis: a prospective cohort of breast cancer patients. Breast Cancer Res. 2007;9:R6.
Early Breast Cancer Trialists’ Collaborative Group (EBCTCG). Tamoxifen for early breast cancer: an overview of the randomised trials. Lancet. 1998;351(9114):1451–67.
Regan MM, Viale G, Mastropasqua MG, et al. Re-evaluating adjuvant breast cancer trials: assessing hormone receptor status by immunohistochemical versus extraction assays. J Natl Cancer Inst. 2006;98(21):1571–81.
Osborne CK. Steroid hormone receptors in breast cancer management. Breast Cancer Res Treat. 1998;51:227–38.
Engel KB, Moore HM. Effects of preanalytical variables on the detection of proteins by immunohistochemistry in formalin-fixed, paraffin-embedded tissue. Arch Pathol Lab Med. 2011;135:537–43.
Goldhirsch A, Glick JH, Gelber RD, et al. Meeting highlights: international expert consensus on the primary therapy of early breast cancer 2005. Ann Oncol. 2005;16(10):1569–83.
Hammond MEH, Hayes DF, Dowsett M, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol. 2010;28(16):2784–95.
American Joint Committee on Cancer. AJCC Cancer Staging Manual, 7th ed. New York: Springer; 2010.
Bloom HJ, Richardson WW. Histological grading and prognosis in breast cancer. Br J Cancer. 1957;11:359–77.
Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow up. Histopathology. 1991;19:403–10.
van Laar R. Design and multiseries validation of a web-based gene expression assay for predicting breast cancer recurrence and patient survival. J Mol Diagn. 2011;13(3):297–304.
Sabatier R, Finetti P, Cervera N, et al. A gene expression signature identifies two prognostic subgroups of basal breast cancer. Breast Cancer Res Treat. 2011;126(2):407–20.
Perou CM, Sorlie T, Eisen MB, et al. Molecular portraits of human breast tumours. Nature. 2000;406(6797):747–52.
Sorlie T, Perou CM, Tibshirania R, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA. 2001;98(19):10869–74.
Kennecke H, Yerushalmi R, Woods R, et al. Metastatic behavior of breast cancer subtypes. J Clin Oncol. 2010;28(20):3271–7.
Raghav KPS, Hernandez-Aya LF, Lei X, et al. Impact of low estrogen/progesterone receptor expression on survival outcomes in breast cancers previously classified as triple negative breast cancers. Cancer. 2012;118(6):1498–506.
Ashworth A. A synthetic lethal therapeutic approach: poly(ADP) ribose polymerase inhibitors for the treatment of cancers deficient in DNA double-strand break repair. J Clin Oncol. 2008;26:3785–90.
Fong PC, Boss DS, Yap TA, et al. Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N Engl J Med. 2009;361(2):123–34.
O’Shaughnessy J, Osborne C, Pippen JE, et al. Iniparib plus chemotherapy in metastatic triple-negative breast cancer. N Engl J Med. 2011;364(3):205–14.
Tutt A, Robson M, Garber JE, et al. Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and advanced breast cancer: a proof-of-concept trial. Lancet. 2010;376(9737):235–44.
Gelmon KA, Tischkowitz M, Mackay H, et al. Olaparib in patients with recurrent high-grade serous or poorly differentiated ovarian carcinoma or triple-negative breast cancer: a phase 2, multicentre, open-label, non-randomised study. Lancet Oncol. 2011;12(9):852–61.
Paik S. Is gene array testing to be considered routine now? Breast. 2011;20(Suppl 3):S87–91.
Ellis MJ, Ding L, Shen, D, et al. Whole-genome analysis informs breast cancer response to aromatase inhibitors. Nature. 2012; 486(7403):353–60.
Carey LA, Perou CM, Livasy CA, et al. Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA. 2006; 295(21): 2492–502.
Acknowledgment
This research was supported by a Grant from the United States Department of Defense (Military Molecular Medicine Initiative MDA W81XWH-05-2-0075, Protocol 01-20006). The opinion and assertions contained herein are the private views of the authors and are not to be construed as official or as representing the views of the Department of the Army or the Department of Defense.
Disclosures
The authors have no conflicts to declare.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Deyarmin, B., Kane, J.L., Valente, A.L. et al. Effect of ASCO/CAP Guidelines for Determining ER Status on Molecular Subtype. Ann Surg Oncol 20, 87–93 (2013). https://doi.org/10.1245/s10434-012-2588-8
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-012-2588-8