Abstract
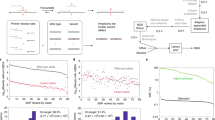
Detection and sequencing of mutations from clinical specimens is often complicated by the presence of an excess of nonmutated cells. To facilitate the detection and sequencing of minority mutations from clinical specimens, we developed wild-type blocking polymerase chain reaction (WTB-PCR). This technique allows sensitive detection of minority mutations in a tissue sample containing excess wild-type DNA. In WTB-PCR, a nonextendable locked nucleic acid (LNA) oligonucleotide binds tightly to a region of wild-type DNA known to develop point mutations. This LNA sequence blocks amplification of wild-type DNA during PCR while permitting amplification of mutant exon 15. Our results show that the LNA blocking oligonucleotide inhibits amplification of wild-type DNA in a dose-dependent manner. WTB-PCR was able to detect mutant DNA in clinical samples of melanoma tissue containing an excess of nonmelanoma cells. This method was also able to detect small amounts of point mutated or tandem mutated DNA diluted with a much larger concentration of wild-type DNA. This rapid and simple assay overcomes the limitations of current methods to detect minority mutations. The potential applications of WTB-PCR include early diagnosis and prognosis of various cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JW, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR and Futreal PA . (2002). Nature, 417, 949–954.
Kaur M, Zhang Y, Liu WH, Tetradis S, Price BD and Makrigiorgos GM . (2002). Mutagenesis, 17, 365–374.
Latorra D, Arar K and Hurley JM . (2003). Mol. Cell Probes, 17, 253–259.
Liu Q, Swiderski P and Sommer SS . (2002). Biotechniques, 33, 129–132, 134–136, 138.
Makrigiorgos GM . (2004). Hum. Mutat., 23, 406–412.
Miller CJ, Cheung M, Sharma A, Clarke L, Helm K, Mauger D and Robertson GP . (2004). J. Invest. Dermatol., 123, 990–992.
Mouritzen P, Nielsen AT, Pfundheller HM, Choleva Y, Kongsbak L and Moller S . (2003). Expert Rev. Mol. Diagn., 3, 27–38.
Oliver DH, Thompson RE, Griffin CA and Eshleman JR . (2000). J. Mol. Diagn., 2, 202–208.
Parsons BL and Heflich RH . (1997). Mutat. Res., 387, 97–121.
SantaLucia Jr J . (1998). Proc. Natl. Acad. Sci. USA, 95, 1460–1465.
Yu D, Mukai M, Liu Q and Steinman CR . (1997). Biotechniques, 23, 714–716, 718–720.
Acknowledgements
We thank Frank Luo for photographs of H&E slides, Marvi Iqbal for extraction of genomic DNA and Raphael Darvish for help in production of GAPDH gels. The work by PLD was supported by funding from the General Clinical Research Center at Harbor-UCLA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dominguez, P., Kolodney, M. Wild-type blocking polymerase chain reaction for detection of single nucleotide minority mutations from clinical specimens. Oncogene 24, 6830–6834 (2005). https://doi.org/10.1038/sj.onc.1208832
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208832
Keywords
This article is cited by
-
Detection of low-frequency mutations in clinical samples by increasing mutation abundance via the excision of wild-type sequences
Nature Biomedical Engineering (2023)
-
LNA blockers for improved amplification selectivity
Scientific Reports (2023)
-
Selective extraction of low-abundance BRAF V600E mutation from plasma, urine, and sputum using ion-tagged oligonucleotides and magnetic ionic liquids
Analytical and Bioanalytical Chemistry (2022)
-
Assessment of the Real-Time PCR Method Claiming to be Specific for Detection and Quantification of the First Commercialised Genome-Edited Plant
Food Analytical Methods (2022)
-
A Comparison Between Full-COLD PCR/HRM and PCR Sequencing for Detection of Mutations in Exon 9 of PIK3CA in Breast Cancer Patients
Applied Biochemistry and Biotechnology (2019)