Abstract
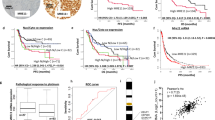
Excision repair cross-complementation group 1 (ERCC1) is a DNA repair enzyme that is frequently defective in non-small cell lung cancer (NSCLC). Although low ERCC1 expression correlates with platinum sensitivity, the clinical effectiveness of platinum therapy is limited, highlighting the need for alternative treatment strategies. To discover new mechanism-based therapeutic strategies for ERCC1-defective tumours, we performed high-throughput drug screens in an isogenic NSCLC model of ERCC1 deficiency and dissected the mechanism underlying ERCC1-selective effects by studying molecular biomarkers of tumour cell response. The high-throughput screens identified multiple clinical poly (ADP-ribose) polymerase 1 and 2 (PARP1/2) inhibitors, such as olaparib (AZD-2281), niraparib (MK-4827) and BMN 673, as being selective for ERCC1 deficiency. We observed that ERCC1-deficient cells displayed a significant delay in double-strand break repair associated with a profound and prolonged G2/M arrest following PARP1/2 inhibitor treatment. Importantly, we found that ERCC1 isoform 202, which has recently been shown to mediate platinum sensitivity, also modulated PARP1/2 sensitivity. A PARP1/2 inhibitor-synthetic lethal siRNA screen revealed that ERCC1 deficiency was epistatic with homologous recombination deficiency. However, ERCC1-deficient cells did not display a defect in RAD51 foci formation, suggesting that ERCC1 might be required to process PARP1/2 inhibitor-induced DNA lesions before DNA strand invasion. PARP1 silencing restored PARP1/2 inhibitor resistance in ERCC1-deficient cells but had no effect in ERCC1-proficient cells, supporting the hypothesis that PARP1 might be required for the ERCC1 selectivity of PARP1/2 inhibitors. This study suggests that PARP1/2 inhibitors as a monotherapy could represent a novel therapeutic strategy for NSCLC patients with ERCC1-deficient tumours.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ . Cancer statistics 2009. Cancer J Clin 2009; 59: 225–249.
Andre F, McShane LM, Michiels S, Ransohoff DF, Altman DG, Reis-Filho JS et al. Biomarker studies: a call for a comprehensive biomarker study registry. Nat Rev Clin Oncol 2011; 8: 171–176.
Curtin NJ . DNA repair dysregulation from cancer driver to therapeutic target. Nat Rev Cancer 2012; 12: 801–817.
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 2004; 350: 2129–2139.
Camidge DR, Bang YJ, Kwak EL, Iafrate AJ, Varella-Garcia M, Fox SB et al. Activity and safety of crizotinib in patients with ALK-positive non-small-cell lung cancer: updated results from a phase 1 study. Lancet Oncol 2012; 13: 1011–1019.
Olaussen KA, Dunant A, Fouret P, Brambilla E, Andre F, Haddad V et al. DNA repair by ERCC1 in non-small-cell lung cancer and cisplatin-based adjuvant chemotherapy. N Engl J Med 2006; 355: 983–991.
Postel-Vinay S, Vanhecke E, Olaussen KA, Lord CJ, Ashworth A, Soria JC . The potential of exploiting DNA-repair defects for optimizing lung cancer treatment. Nat Rev Clin Oncol 2012; 9: 144–155.
Simon GR, Schell MJ, Begum M, Kim J, Chiappori A, Haura E et al. Preliminary indication of survival benefit from ERCC1 and RRM1-tailored chemotherapy in patients with advanced nonsmall cell lung cancer: evidence from an individual patient analysis. Cancer 2012; 118: 2525–2531.
Vilmar AC, Santoni-Rugiu E, Sorensen JB . ERCC1 and histopathology in advanced NSCLC patients randomized in a large multicenter phase III trial. Ann Oncol 2010; 21: 1817–1824.
Fagbemi AF, Orelli B, Scharer OD . Regulation of endonuclease activity in human nucleotide excision repair. DNA Repair 2011; 10: 722–729.
Kirschner K, Melton DW . Multiple roles of the ERCC1-XPF endonuclease in DNA repair and resistance to anticancer drugs. Anticancer Res 2010; 30: 3223–3232.
Tripsianes K, Folkers G, Ab E, Das D, Odijk H, Jaspers NG et al. The structure of the human ERCC1/XPF interaction domains reveals a complementary role for the two proteins in nucleotide excision repair. Structure 2005; 13: 1849–1858.
Bergstralh DT, Sekelsky J . Interstrand crosslink repair: can XPF-ERCC1 be let off the hook? Trends Genet 2008; 24: 70–76.
Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 2005; 434: 917–921.
Friboulet L, Olaussen KA, Pignon JP, Shepherd FA, Tsao M, Graziano S et al. ERCC1 isoform expression and DNA repair in non-small cell lung cancer. N Engl J Med 2013; 368: 1101–1110.
Zhang YW, Regairaz M, Seiler JA, Agama KK, Doroshow JH, Pommier Y . Poly(ADP-ribose) polymerase and XPF-ERCC1 participate in distinct pathways for the repair of topoisomerase I-induced DNA damage in mammalian cells. Nucleic Acids Res 2011; 39: 3607–3620.
Wang B, Chu D, Feng Y, Xin Y, Myers P, Post L et al. Novel PARP inhibitors with potent antitumor activity as single-agent and combination therapies. Mol Cancer Ther 2009; 8 (Suppl 12): A121.
Patterson M, Murray J, Curtin NJ . Stability of PARP inhibition by BMN 673 in human PBMCs and Leukaemic Cell Cultures. Eur J Cancer 2012; 48 (Suppl 6): 106 (Abstract 348).
Turner NC, Lord CJ, Iorns E, Brough R, Swift S, Elliott R et al. A synthetic lethal siRNA screen identifying genes mediating sensitivity to a PARP inhibitor. Embo J 2008; 27: 1368–1377.
Murai J, Huang SY, Das BB, Renaud A, Zhang Y, Doroshow JH et al. Trapping of PARP1 and PARP2 by Clinical PARP Inhibitors. Cancer Res 2012; 72: 5588–5599.
Kedar PS, Stefanick DF, Horton JK, Wilson SH . Increased PARP-1 association with DNA in alkylation damaged, PARP-inhibited mouse fibroblasts. Mol Cancer Res 2012; 10: 360–368.
Yin M, Yan J, Martinez-Balibrea E, Graziano F, Lenz HJ, Kim HJ et al. ERCC1 and ERCC2 polymorphisms predict clinical outcomes of oxaliplatin-based chemotherapies in gastric and colorectal cancer: a systemic review and meta-analysis. Clin Cancer Res 2011; 17: 1632–1640.
Langer CJ . Exploring biomarkers in head and neck cancer. Cancer 2012; 118: 3882–3892.
Vilmar AC, Sorensen JB . Customising chemotherapy in advanced nonsmall cell lung cancer: daily practice and perspectives. Eur Respir Rev 2011; 20: 45–52.
Metzger R, Bollschweiler E, Holscher AH, Warnecke-Eberz U . ERCC1: impact in multimodality treatment of upper gastrointestinal cancer. Future Oncol 2010; 6: 1735–1749.
Steffensen KD, Waldstrom M, Jakobsen A . The relationship of platinum resistance and ERCC1 protein expression in epithelial ovarian cancer. Int J Gynecol Cancer 2009; 19: 820–825.
Chen S, Zhang J, Wang R, Luo X, Chen H . The platinum-based treatments for advanced non-small cell lung cancer, is low/negative ERCC1 expression better than high/positive ERCC1 expression? A meta-analysis. Lung cancer 2010; 70: 63–70.
Lord RV, Brabender J, Gandara D, Alberola V, Camps C, Domine M et al. Low ERCC1 expression correlates with prolonged survival after cisplatin plus gemcitabine chemotherapy in non-small cell lung cancer. Clin Cancer Res 2002; 8: 2286–2291.
Rehman FL, Lord CJ, Ashworth A . Synthetic lethal approaches to breast cancer therapy. Nat Rev Clin Oncol 2010; 7: 718–724.
Johnson N, Li YC, Walton ZE, Cheng KA, Li D, Rodig SJ et al. Compromised CDK1 activity sensitizes BRCA-proficient cancers to PARP inhibition. Nat Med 2011; 17: 875–882.
Sijbers AM, van der Spek PJ, Odijk H, van den Berg J, van Duin M, Westerveld A et al. Mutational analysis of the human nucleotide excision repair gene ERCC1. Nucleic Acids Res 1996; 24: 3370–3380.
Dabholkar M, Vionnet J, Parker R, Bostickbruton F, Dobbins A, Reed E . Expression of an alternatively spliced ercc1 messenger-RNA species, is related to reduced DNA-repair efficiency in human T-lymphocytes. Oncol Rep 1995; 2: 209–214.
Sun Y, Li T, Ma K, Tian Z, Zhu Y, Chen F et al. The impacts of ERCC1 gene exon VIII alternative splicing on cisplatin-resistance in ovarian cancer cells. Cancer Invest 2009; 27: 891–897.
Cheng H, Zhang Z, Borczuk A, Powell CA, Balajee AS, Lieberman HB et al. PARP inhibition selectively increases sensitivity to cisplatin in ERCC1-low non-small cell lung cancer cells. Carcinogenesis 2013; 34: 739–749.
Niedernhofer LJ, Essers J, Weeda G, Beverloo B, de Wit J, Muijtjens M et al. The structure-specific endonuclease Ercc1-Xpf is required for targeted gene replacement in embryonic stem cells. Embo J 2001; 20: 6540–6549.
Langelier MF, Planck JL, Roy S, Pascal JM . Crystal structures of poly(ADP-ribose) polymerase-1 (PARP-1) zinc fingers bound to DNA: structural and functional insights into DNA-dependent PARP-1 activity. J Biol Chem 2011; 286: 10690–10701.
Langelier MF, Planck JL, Roy S, Pascal JM . Structural basis for DNA damage-dependent poly(ADP-ribosyl)ation by human PARP-1. Science 2012; 336: 728–732.
Bajrami I, Kigozi A, Van Weverwijk A, Brough R, Frankum J, Lord CJ et al. Synthetic lethality of PARP and NAMPT inhibition in triple-negative breast cancer cells. EMBO Mol Med 2012; 4: 1087–1096.
Graeser M, McCarthy A, Lord CJ, Savage K, Hills M, Salter J et al. A marker of homologous recombination predicts pathologic complete response to neoadjuvant chemotherapy in primary breast cancer. Clin Cancer Res 2010; 16: 6159–6168.
Edwards SL, Brough R, Lord CJ, Natrajan R, Vatcheva R, Levine DA et al. Resistance to therapy caused by intragenic deletion in BRCA2. Nature 2008; 451: 1111–1115.
Lord CJ, Martin SA, Ashworth A . RNA interference screening demystified. J Clin Pathol 2009; 62: 195–200.
Acknowledgements
We thank David Roberston and Marieke Aarts for assistance with confocal microscopy and FACS analysis, and Jerry Shen and Len Post at Biomarin for the provision of BMN 673. We also acknowledge NHS funding to the NIHR Royal Marsden Hospital BRC. This work was supported by grants from Cancer Research UK and The European Union as part of the FP7 teams ‘DDResponse’ and ‘Eurocan’. SPV is supported by a Translational Research Fellowship from the European Society of Medical Oncology (2011 and 2012) and by funding from the Institut National du Cancer: Bourse pour la Formation à la Recherche Translationnelle (2011)
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
AA and CJL are inventors on patents describing the use of PARP inhibitors and may benefit from the ICR ‘Rewards to inventors scheme’. All other authors declare no conflicts of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Postel-Vinay, S., Bajrami, I., Friboulet, L. et al. A high-throughput screen identifies PARP1/2 inhibitors as a potential therapy for ERCC1-deficient non-small cell lung cancer. Oncogene 32, 5377–5387 (2013). https://doi.org/10.1038/onc.2013.311
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2013.311
Keywords
This article is cited by
-
Olaparib maintenance versus placebo in platinum-sensitive non-small cell lung cancer: the Phase 2 randomized PIPSeN trial
British Journal of Cancer (2024)
-
Circulating tumor cell copy-number heterogeneity in ALK-rearranged non-small-cell lung cancer resistant to ALK inhibitors
npj Precision Oncology (2021)
-
Optimal control nodes in disease-perturbed networks as targets for combination therapy
Nature Communications (2019)
-
Synthetically Lethal BMN 673 (Talazoparib) Loaded Solid Lipid Nanoparticles for BRCA1 Mutant Triple Negative Breast Cancer
Pharmaceutical Research (2018)
-
ERCC1 assessment in upfront treatment with and without cisplatin-based chemotherapy in stage IIIB/IV non-squamous non-small cell lung cancer
Medical Oncology (2018)