Abstract
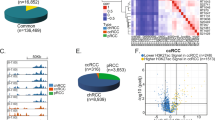
The detection of promoter region hypermethylation and transcriptional silencing has facilitated the identification of candidate renal cell carcinoma (RCC) tumour suppressor genes (TSGs). We have used a genome-wide strategy (methylated DNA immunoprecipitation (MeDIP) and whole-genome array analysis in combination with high-density expression array analysis) to identify genes that are frequently methylated and silenced in RCC. MeDIP analysis on 9 RCC tumours and 3 non-malignant normal kidney tissue samples was performed, and an initial shortlist of 56 candidate genes that were methylated by array analysis was further investigated; 9 genes were confirmed to show frequent promoter region methylation in primary RCC tumour samples (KLHL35 (39%), QPCT (19%), SCUBE3 (19%), ZSCAN18 (32%), CCDC8 (35%), FBN2 (34%), ATP5G2 (36%), PCDH8 (58%) and CORO6 (22%)). RNAi knockdown for KLHL35, QPCT, SCUBE3, ZSCAN18, CCDC8 and FBN2 resulted in an anchorage-independent growth advantage. Tumour methylation of SCUBE3 was associated with a significantly increased risk of cancer death or relapse (P=0.0046). The identification of candidate epigenetically inactivated RCC TSGs provides new insights into renal tumourigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Annes JP, Munger JS, Rifkin DB . (2003). Making sense of latent TGFbeta activation. J Cell Sci 116: 217–224.
Battagli C, Uzzo RG, Dulaimi E, Ibanez de Caceres I, Krassenstein R, Al-Saleem T et al. (2003). Promoter hypermethylation of tumor suppressor genes in urine from kidney cancer patients. Cancer Res 63: 8695–8699.
Bray F, Sankila R, Ferlay J, Parkin DM . (2002). Estimates of cancer incidence and mortality in Europe in 1995. Eur J Cancer 38: 99–166.
Breault JE, Shiina H, Igawa M, Ribeiro-Filho LA, Deguchi M, Enokida H et al. (2005). Methylation of the gamma-catenin gene is associated with poor prognosis of renal cell carcinoma. Clin Cancer Res 11: 557–564.
Chaudhry SS, Cain SA, Morgan A, Dallas SL, Shuttleworth CA, Kielty CM . (2007). Fibrillin-1 regulates the bioavailability of TGFbeta1. J Cell Biol 176: 355–367.
Chen H, Suzuki M, Nakamura Y, Ohira M, Ando S, Iida T et al. (2005). Aberrant methylation of FBN2 in human non-small cell lung cancer. Lung Cancer 50: 43–49.
Chowdhury S, Larkin JM, Gore ME . (2008). Recent advances in the treatment of renal cell carcinoma and the role of targeted therapies. Eur J Cancer 44: 2152–2161.
Christoph F, Weikert S, Kempkensteffen C, Krause H, Schostak M, Köllermann J et al. (2006). Promoter hypermethylation profile of kidney cancer with new proapoptotic p53 target genes and clinical implications. Clin Cancer Res 12: 5040–5046.
Clifford SC, Prowse AH, Affara NA, Buys CH, Maher ER . (1998). Inactivation of the von Hippel-Lindau (VHL) tumour suppressor gene and allelic losses at chromosome arm 3p in primary renal cell carcinoma: evidence for a VHL-independent pathway in clear cell renal tumourigenesis. Genes Chromosomes Cancer 22: 200–209.
Costa LJ, Drabkin HA . (2007). Renal cell carcinoma: new developments in molecular biology and potential for targeted therapies. Oncologist 12: 1404–1415.
Dahl E, Wiesmann F, Woenckhaus M, Stoehr R, Wild PJ, Veeck J et al. (2007). Frequent loss of SFRP1 expression in multiple human solid tumours: association with aberrant promoter methylation in renal cell carcinoma. Oncogene 26: 5680–5691.
Dalgliesh GL, Furge K, Greenman C, Chen L, Bignell G, Butler A et al. (2010). Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463: 360–363.
Dyer MR, Walker JE . (1993). Sequences of members of the human gene family for the c subunit of mitochondrial ATP synthase. Biochem J 293 (Part 1): 51–64.
Fischer WH, Spiess J . (1987). Identification of a mammalian glutaminyl cyclase converting glutaminyl into pyroglutamyl peptides. Proc Natl Acad Sci USA 84: 3628–3632.
Foster K, Prowse A, van den Berg A, Fleming S, Hulsbeek MM, Crossey PA et al. (1994). Somatic mutations of the von Hippel-Lindau disease tumour suppressor gene in non-familial clear cell renal carcinoma. Hum Mol Genet 3: 2169–2173.
Grimmond S, Larder R, Van Hateren N, Siggers P, Hulsebos TJ, Arkell R et al. (2000). Cloning, mapping, and expression analysis of a gene encoding a novel mammalian EGF-related protein (SCUBE1). Genomics 70: 74–81.
Herman JG, Latif F, Weng Y, Lerman MI, Zbar B, Liu S et al. (1994). Silencing of the VHL tumor-suppressor gene by DNA methylation in renal carcinoma. Proc Natl Acad Sci USA 91: 9700–9704.
Hogg RP, Honorio S, Martinez A, Agathanggelou A, Dallol A, Fullwood P et al. (2002). Frequent 3p allele loss and epigenetic inactivation of the RASSF1A tumour suppressor gene from region 3p21.3 in head and neck squamous cell carcinoma. Eur J Cancer 38: 1585–1592.
Hoque MO, Begum S, Topaloglu O, Chatterjee A, Rosenbaum E, Van Criekinge W et al. (2006). Quantitation of promoter methylation of multiple genes in urine DNA and bladder cancer detection. J Natl Cancer Inst 98: 996–1004.
Hüttemann M, Lee I, Pecinova A, Pecina P, Przyklenk K, Doan JW . (2008). Regulation of oxidative phosphorylation, the mitochondrial membrane potential, and their role in human disease. J Bioenerg Biomembr 40: 445–456.
Ibanez de Caceres I, Dulaimi E, Hoffman AM, Al-Saleem T, Uzzo RG, Cairns P . (2006). Identification of novel target genes by an epigenetic reactivation screen of renal cancer. Cancer Res 66: 5021–5028.
Imoto I, Izumi H, Yokoi S, Hosoda H, Shibata T, Hosoda F et al. (2006). Frequent silencing of the candidate tumor suppressor PCDH20 by epigenetic mechanism in non-small-cell lung cancers. Cancer Res 66: 4617–4626.
Latif F, Tory K, Gnarra J, Yao M, Duh FM, Orcutt ML et al. (1993). Identification of the von Hippel-Lindau disease tumor suppressor gene. Science 260: 1317–1320.
Leshchenko VV, Kuo P-Y, Shaknovich R, Yang DT, Gellen T, Petrich A et al. (2010). Genome wide DNA methylation analysis reveals novel targets for drug development in mantle cell lymphoma. Blood 116: 1025–1034.
Li B, Carey M, Workman JL . (2007). The role of chromatin during transcription. Cell 128: 707–719.
Lodygin D, Epanchintsev A, Menssen A, Diebold J, Hermeking H . (2005). Functional epigenomics identifies genes frequently silenced in prostate cancer. Cancer Res 65: 4218–4227.
Maxwell PH, Wiesener MS, Chang GW, Clifford SC, Vaux EC, Cockman ME et al. (1999). The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399: 271–275.
McRonald FE, Morris MR, Gentle D, Winchester L, Baban D, Ragoussis J et al. (2009). CpG methylation profiling in VHL related and VHL unrelated renal cell carcinoma. Mol Cancer 8: 31.
Morris MR, Gentle D, Abdulrahman M, Clarke N, Brown M, Kishida T et al. (2008). Functional epigenomics approach to identify methylated candidate tumour suppressor genes in renal cell carcinoma. Br J Cancer 98: 496–501.
Morris MR, Gentle D, Abdulrahman M, Maina EN, Gupta K, Banks RE et al. (2005). Tumor suppressor activity and epigenetic inactivation of hepatocyte growth factor activator inhibitor type 2/SPINT2 in papillary and clear cell renal cell carcinoma. Cancer Res 65: 4598–4606.
Morris MR, Hesson LB, Wagner KJ, Morgan NV, Astuti D, Lees RD et al. (2003). Multigene methylation analysis of Wilms′ tumour and adult renal cell carcinoma. Oncogene 22: 6794–6801.
Morris MR, Maina E, Morgan NV, Gentle D, Astuti D, Moch H et al. (2004). Molecular genetic analysis of FIH-1, FH, and SDHB candidate tumour suppressor genes in renal cell carcinoma. J Clin Pathol 57: 706–711.
Morris MR, Ricketts C, Gentle D, Abdulrahman M, Clarke N, Brown M et al. (2010). Identification of candidate tumour suppressor genes frequently methylated in renal cell carcinoma. Oncogene 29: 2104–2117.
Morrissey C, Martinez A, Zatyka M, Agathanggelou A, Honorio S, Astuti D et al. (2001). Epigenetic inactivation of the RASSF1A 3p21.3 tumor suppressor gene in both clear cell and papillary renal cell carcinoma. Cancer Res 61: 7277–7281.
Muthusamy V, Duraisamy S, Bradbury CM, Hobbs C, Curley DP, Nelson B et al. (2006). Epigenetic silencing of novel tumor suppressors in malignant melanoma. Cancer Res 66: 11187–11193.
Ohtani-Fujita N, Fujita T, Aoike A, Osifchin NE, Robbins PD, Sakai T . (1993). CpG methylation inactivates the promoter activity of the human retinoblastoma tumor-suppressor gene. Oncogene 8: 1063–1067.
Pflug BR, Zheng H, Udan MS, D'Antonio JM, Marshall FF, Brooks JD et al. (2007). Endothelin-1 promotes cell survival in renal cell carcinoma through the ET(A) receptor. Cancer Lett 246: 139–148.
Pohl T, Zimmer M, Mugele K, Spiess J . (1991). Primary structure and functional expression of a glutaminyl cyclase. Proc Natl Acad Sci USA 88: 10059–10063.
Ricketts C, Woodward ER, Killick P, Morris MR, Astuti D, Latif F et al. (2008). Germline SDHB mutations and familial renal cell carcinoma. J Natl Cancer Inst 100: 1260–1262.
Roadcap DW, Clemen CS, Bear JE . (2008). The role of mammalian coronins in development and disease. Subcell Biochem 48: 124–135.
Robinson PN, Godfrey M . (2000). The molecular genetics of Marfan syndrome and related microfibrillopathies. J Med Genet 37: 9–25.
Sato N, Fukushima N, Maitra A, Matsubayashi H, Yeo CJ, Cameron JL et al. (2003). Discovery of novel targets for aberrant methylation in pancreatic carcinoma using high-throughput microarrays. Cancer Res 63: 3735–3742.
Thouënnon E, Elkahloun AG, Guillemot J, Gimenez-Roqueplo A-P, Bertherat J, Pierre A et al. (2007). Identification of potential gene markers and insights into the pathophysiology of pheochromocytoma malignancy. J Clin Endocrinol Metab 92: 4865–4872.
Tsunoda S, Smith E, De Young NJ, Wang X, Tian Z-Q, Liu J-F et al. (2009). Methylation of CLDN6, FBN2, RBP1, RBP4, TFPI2, and TMEFF2 in esophageal squamous cell carcinoma. Oncol Rep 21: 1067–1073.
Urakami S, Shiina H, Enokida H, Hirata H, Kawamoto K, Kawakami T et al. (2006). Wnt antagonist family genes as biomarkers for diagnosis, staging, and prognosis of renal cell carcinoma using tumor and serum DNA. Clin Cancer Res 12: 6989–6997.
Yagi K, Akagi K, Hayashi H, Nagae G, Tsuji S, Isagawa T et al. (2010). Three DNA methylation epigenotypes in human colorectal cancer. Clin Cancer Res 16: 21–33.
Yamada D, Kikuchi S, Williams YN, Sakurai-Yageta M, Masuda M, Maruyama T et al. (2006). Promoter hypermethylation of the potential tumor suppressor DAL-1/4.1B gene in renal clear cell carcinoma. Int J Cancer 118: 916–923.
Yamashita K, Upadhyay S, Osada M, Hoque MO, Xiao Y, Mori M et al. (2002). Pharmacologic unmasking of epigenetically silenced tumor suppressor genes in esophageal squamous cell carcinoma. Cancer Cell 2: 485–495.
Ying J, Li H, Seng TJ, Langford C, Srivastava G, Tsao SW et al. (2006). Functional epigenetics identifies a protocadherin PCDH10 as a candidate tumor suppressor for nasopharyngeal, esophageal and multiple other carcinomas with frequent methylation. Oncogene 25: 1070–1080.
Yu JS, Koujak S, Nagase S, Li C-M, Su T, Wang X et al. (2008). PCDH8, the human homolog of PAPC, is a candidate tumor suppressor of breast cancer. Oncogene 27: 4657–4665.
Zhang H, Apfelroth SD, Hu W, Davis EC, Sanguineti C, Bonadio J et al. (1994). Structure and expression of fibrillin-2, a novel microfibrillar component preferentially located in elastic matrices. J Cell Biol 124: 855–863.
Acknowledgements
We thank Cancer Research UK for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Morris, M., Ricketts, C., Gentle, D. et al. Genome-wide methylation analysis identifies epigenetically inactivated candidate tumour suppressor genes in renal cell carcinoma. Oncogene 30, 1390–1401 (2011). https://doi.org/10.1038/onc.2010.525
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.525
Keywords
This article is cited by
-
Analysis and experimental validation of necroptosis-related molecular classification, immune signature and feature genes in Alzheimer’s disease
Apoptosis (2024)
-
Inactivation of ZSCAN18 by promoter hypermethylation drives the proliferation via attenuating TP53INP2-mediated autophagy in gastric cancer cells
Clinical Epigenetics (2023)
-
Identification and validation of a DNA methylation-driven gene-based prognostic model for clear cell renal cell carcinoma
BMC Genomics (2023)
-
Predicting congenital renal tract malformation genes using machine learning
Scientific Reports (2023)
-
DNA methylation-mediated low expression of ZNF582 promotes the proliferation, migration, and invasion of clear cell renal cell carcinoma
Clinical and Experimental Nephrology (2023)