Abstract
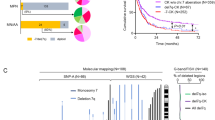
Historically, genes targeted by recurrent chromosomal deletions have been identified within the smallest genomic region shared in all patients, the minimally deleted region (MDR). However, deletions this small do not occur in all patients and are a simplification of the impact larger heterogeneous deletions have during carcinogenesis. We use the example of 13q14 deletions in chronic lymphocytic leukemia to show that genes outside MDRs are associated with disease progression. Genomic profiling of 224 patients identified 205 copy number alterations on chromosome 13 in 132 cases. Deletions including DLEU2 were heterogeneous (845 Kb–96.2 Mb) and identified two breakpoint cluster regions within short interspersed nuclear elements proximal to DLEU2 and within long interspersed nuclear elements/L1 repeats distal to GUCY1B2. After defining a deletion class on the basis of size and location, we show that (a) at diagnosis, larger deletions (class II) were associated with a significantly increased risk of disease progression (odds ratio=12.3; P=0.005), (b) in progressive patients, class II deletions were enriched (P=0.02) and (c) this association was independent of IgVH mutational status, ZAP70 expression and ATM/TP53 deletion. Deletion of a 1 Mb gene cluster (48.2–49.2 Mb), including SETDB2, PHF11 and RCBTB1, was significantly associated (P<0.01) with disease progression. Here, we show that the deletion of genes outside MDRs can influence clinical outcome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Mitelman F, Mertens F, Johansson B . A breakpoint map of recurrent chromosomal rearrangements in human neoplasia. Nat Genet 1997; 15: 417–474.
Fitchett M, Griffiths MJ, Oscier DG, Johnson S, Seabright M . Chromosome abnormalities involving band 13q14 in hematologic malignancies. Cancer Genet Cytogenet 1987; 24: 143–150.
Liu Y, Grandér D, Söderhäll S, Juliusson G, Gahrton G, Einhorn S . Retinoblastoma gene deletions in B-cell chronic lymphocytic leukemia. Genes Chromosomes Cancer 1992; 4: 250–256.
Liu Y, Szekely L, Grandér D, Söderhäll S, Juliusson G, Gahrton G et al. Chronic lymphocytic leukemia cells with allelic deletions at 13q14 commonly have one intact RB1 gene: evidence for a role of an adjacent locus. Proc Natl Acad Sci USA 1993; 90: 8697–8701.
Stilgenbauer S, Döhner H, Bulgay-Mörschel M, Weitz S, Bentz M, Lichter P . High frequency of monoallelic retinoblastoma gene deletion in B-cell chronic lymphoid leukemia shown by interphase cytogenetics. Blood 1993; 81: 2118–2124.
Kalachikov S, Migliazza A, Cayanis E, Fracchiolla NS, Bonaldo MF, Lawton L et al. Cloning and gene mapping of the chromosome 13q14 region deleted in chronic lymphocytic leukemia. Genomics 1997; 42: 369–377.
Bullrich F, Veronese ML, Kitada S, Jurlander J, Caligiuri MA, Reed JC et al. Minimal region of loss at 13q14 in B-cell chronic lymphocytic leukemia. Blood 1996; 88: 3109–3115.
Stilgenbauer S, Nickolenko J, Wilhelm J, Wolf S, Weitz S, Döhner K et al. Expressed sequences as candidates for a novel tumor suppressor gene at band 13q14 in B-cell chronic lymphocytic leukemia and mantle cell lymphoma. Oncogene 1998; 16: 1891–1897.
Liu Y, Corcoran M, Rasool O, Ivanova G, Ibbotson R, Grandér D et al. Cloning of two candidate tumor suppressor genes within a 10 kb region on chromosome 13q14, frequently deleted in chronic lymphocytic leukemia. Oncogene 1997; 15: 2463–2473.
Migliazza A, Bosch F, Komatsu H, Cayanis E, Martinotti S, Toniato E et al. Nucleotide sequence, transcription map, and mutation analysis of the 13q14 chromosomal region deleted in B-cell chronic lymphocytic leukemia. Blood 2001; 97: 2098–2104.
Bullrich F, Fujii H, Calin G, Mabuchi H, Negrini M, Pekarsky Y et al. Characterization of the 13q14 tumor suppressor locus in CLL: identification of ALT1, an alternative splice variant of the LEU2 gene. Cancer Res 2001; 61: 6640–6648.
Mertens D, Wolf S, Schroeter P, Schaffner C, Döhner H, Stilgenbauer S et al. Down-regulation of candidate tumor suppressor genes within chromosome band 13q14.3 is independent of the DNA methylation pattern in B-cell chronic lymphocytic leukemia. Blood 2002; 99: 4116–4121.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99: 15524–15529.
Klein U, Lia M, Crespo M, Siegel R, Shen Q, Mo T et al. The DLEU2/miR-15a/16–1 cluster controls B cell proliferation and its deletion leads to chronic lymphocytic leukemia. Cancer Cell 2010; 17: 28–40.
Lerner M, Corcoran M, Cepeda D, Nielsen ML, Zubarev R, Pontén F et al. The RBCC gene RFP2 (Leu5) encodes a novel transmembrane E3 ubiquitin ligase involved in ERAD. Mol Biol Cell 2007; 18: 1670–1682.
Palamarchuk A, Efanov A, Nazaryan N, Santanam U, Alder H, Rassenti L et al. 13q14 deletions in CLL involve cooperating tumor suppressors. Blood 2010; 115: 3916–3922.
Ouillette P, Erba H, Kujawski L, Kaminski M, Shedden K, Malek SN . Integrated genomic profiling of chronic lymphocytic leukemia identifies subtypes of deletion 13q14. Cancer Res 2008; 68: 1012–1021.
Ouillette P, Fossum S, Parkin B, Ding L, Bockenstedt P, Al-Zoubi A et al. Aggressive chronic lymphocytic leukemia with elevated genomic complexity is associated with multiple gene defects in the response to DNA double-strand breaks. Clin Cancer Res 2010; 16: 835–847.
Catovsky D, Richards S, Matutes E, Oscier D, Dyer MJ, Bezares RF et al. Assessment of fludarabine plus cyclophosphamide for patients with chronic lymphocytic leukaemia (the LRF CLL4 Trial): a randomised controlled trial. Lancet 2007; 370: 230–239.
ISCN. An International System for Human Cytogenetic Nomenclature: Recommendations of the International Standing Committee on Human Cytogenetic Nomenclature. S.Karger: Basel, 2009.
Best OG, Ibbotson RE, Parker AE, Davis ZA, Orchard JA, Oscier DG . ZAP-70 by flow cytometry: a comparison of different antibodies, anticoagulants, and methods of analysis. Cytometry B Clin Cytom 2006; 70: 235–241.
Oscier DG, Gardiner AC, Mould SJ, Glide S, Davis ZA, Ibbotson RE et al. Multivariate analysis of prognostic factors in CLL: clinical stage, IGVH gene mutational status, and loss or mutation of the p53 gene are independent prognostic factors. Blood 2002; 100: 1177–1184.
Hamblin TJ, Davis Z, Gardiner A, Oscier DG, Stevenson FK . Unmutated Ig V(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood 1999; 94: 1848–1854.
Dupont WD, Plummer WD . Power and sample size calculations: a review and computer program. Controlled Clinical Trials 1998; 19: 589–601.
Pfeifer D, Pantic M, Skatulla I, Rawluk J, Kreutz C, Martens UM et al. Genome-wide analysis of DNA copy number changes and LOH in CLL using high-density SNP arrays. Blood 2007; 109: 1202–1210.
Lerner M, Harada M, Loven J, Castro J, Davis Z, Oscier D et al. DLEU2 frequently deleted in malignancy, functions as a critical host gene of the cell cycle inhibitory microRNA's miR15a and miR 16–1. Exp Cell Res 2009; 315: 2941–52.
Falandry C, Fourel G, Galy V, Ristriani T, Horard B, Bensimon E et al. CLLD8/KMT1F is a lysine methyltrasferase that is important for chromosome segregation. J Biol Chem 2010; 285: 20234–20241.
Chuikov S, Kurash JK, Wilson JR, Xiao B, Justin N, Ivanov GS et al. Regulation of p53 activity through lysine methylation. Nature 2004; 432: 353–360.
Clarke E, Rahman N, Page N, Rolph MS, Stewart GJ, Jones GJ . Functional characterization of the atopy-associated gene PHF11. J Allergy Clin Immunol 2008; 121: 1148–1154.
Plafker KS, Singer JD, Plafker SM . The ubiquitin conjugating enzyme, UbcM2, engages in novel interactions with components of cullin-3 based E3 ligases. Biochemistry 2009; 48: 3527–3537.
Oscier D, Wade R, Davis Z, Morilla A, Best G, Richards S et al. Prognostic factors identified three risk groups in the LRF CLL4 trial, independent of treatment allocation. Haematologica 2010; 95: 1705–1712.
Acknowledgements
This study was funded by Leukaemia and Lymphoma Research and Cancer Research UK. The LRF CLL4 trial was funded by a core grant from Leukaemia and Lymphoma Research, with associated research work supported by the MRC (G8223452) and Cancer Research UK, and laboratory studies by the Arbib Foundation, Schering Healthcare UK, Schering AG—Germany and Leukaemia and Lymphoma Research. The authors gratefully acknowledge all patients and clinicians who participated in the trial. We would like to thank Professor Nick Cross for help with the preparation of this manuscript.
Author contributions
This work was funded by grants awarded to JCS; HP, TC and BDY performed the genomic profiling experiments; MG coordinated the array-CGH profiling; HP, MJRZ and JCS performed the microarray analysis. AG and AP performed the cytogenetic and molecular diagnostic assays. MJRZ, RW, AP and AC conducted statistical analyses; DGO contributed patient samples and data; DGO and JCS initiated and designed the study; JCS wrote the manuscript with contributions from HP, MJRZ and DGO; and all authors critically reviewed the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Parker, H., Rose-Zerilli, M., Parker, A. et al. 13q deletion anatomy and disease progression in patients with chronic lymphocytic leukemia. Leukemia 25, 489–497 (2011). https://doi.org/10.1038/leu.2010.288
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2010.288
Keywords
This article is cited by
-
Effects of Chemotherapy on the Immune System: Implications for Cancer Treatment and Patient Outcomes
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Which prognostic marker is responsible for the clinical heterogeneity in CLL with 13q deletion?
Molecular Cytogenetics (2021)
-
Emerging roles of H3K9me3, SETDB1 and SETDB2 in therapy-induced cellular reprogramming
Clinical Epigenetics (2019)
-
Combining cytogenetic and epigenetic approaches in chronic lymphocytic leukemia improves prognosis prediction for patients with isolated 13q deletion
Clinical Epigenetics (2017)
-
CALR mutational status identifies different disease subtypes of essential thrombocythemia showing distinct expression profiles
Blood Cancer Journal (2017)