Abstract
Noonan syndrome (NS) is characterized by short stature, characteristic facial features, and heart defects. Recently, missense mutations of PTPN11, the gene encoding protein tyrosine phosphatase (PTP) SHP-2, were identified in patients with NS. Further, somatic mutations in PTPN11 were detected in childhood leukemia. Recent studies showed that the phosphatase activities of five mutations identified in NS and juvenile myelomonocytic leukemia (JMML) were increased. However, the functional properties of the other mutations remain unidentified. In this study, in order to clarify the differences between the mutations identified in NS and leukemia, we examined the phosphatase activity of 14 mutants of SHP-2. We identified nine mutations, including a novel F71I mutation, in 16 of 41 NS patients and two mutations, including a novel G503V mutation, in three of 29 patients with leukemia. Immune complex phosphatase assays of individual mutants transfected in COS7 cells showed that ten mutants identified in NS and four mutants in leukemia showed 1.4-fold to 12.7-fold increased activation compared with wild-type SHP-2. These results suggest that the pathogenesis of NS and leukemia is associated with enhanced phosphatase activity of mutant SHP-2. A comparison of the phosphatase activity in each mutant and a review of previously reported cases showed that high phosphatase activity observed in mutations at codons 61, 71, 72, and 76 was significantly associated with leukemogenesis.
Similar content being viewed by others
Introduction
Noonan syndrome (NS; MIM#163950) is an autosomal dominant developmental disorder characterized by facial dysmorphism including hypertelorism, low-set ears and ptosis, short stature, skeletal abnormalities, and heart defects (Mendez and Opitz 1985; Allanson 1987). Frequently observed features in NS patients are pulmonary stenosis, hypertrophic cardiomyopathy, chest deformities, webbed and short neck, mental retardation, and genitourinary defects including cryptorchidism in males, and bleeding diathesis due to factor XI deficiency (Mendez and Opitz 1985). The incidence of this syndrome is estimated to be 1:1,000 to 1:2,500 live births (Allanson 1987). Tartaglia et al. (2001) have identified missense mutations in PTPN11, a gene encoding tyrosine phosphatase SHP-2, in 45% of clinically diagnosed NS patients. Patients with NS have been known to be associated with juvenile myelomonocytic leukemia (JMML), a myeloproliferative disorder characterized by excessive production of myelomonocytic cells (Tartaglia et al. 2003). Interestingly, somatic mutations in PTPN11 were identified in 34% of JMML cells and in a small percentage of patients with myelodysplastic syndrome (MDS), acute myeloid leukemia (AML), and acute lymphoid leukemia (ALL) from non-NS-leukemic patients (Tartaglia et al. 2003; Loh et al. 2004; Tartaglia et al. 2004).
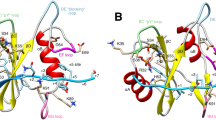
SHP-2, a widely expressed cytoplasmic tyrosine phosphatase, has been implicated in signal transduction pathways elicited by growth factors, cytokines, hormones, and extracellular matrix (Servidei et al. 1998; Feng 1999; Neel et al. 2003). One of the well-known pathways is the activation of the RAS/MAPK (mitogen-activated protein kinase) cascade induced by epidermal, fibroblast, and hepatocyte growth factors (Feng 1999; Neel et al. 2003). SHP-2 contains a tandem array of two SH2 domains at its N terminus, a catalytic domain in the middle, and a C-terminal domain that contains tyrosine phosphorylation sites. A crystallographic analysis indicates that intramolecular conformational change controls its catalytic activity (Hof et al. 1998). In the inactive state, the N-SH2 domain directly binds the PTP domain and blocks its catalytic activity. Once the N-SH2 domain binds phosphotyrosine peptide, conformational change results in the active state. Previously identified mutations in NS and leukemic patients were primarily restricted in or around the interacting face of the N-SH2 and PTP domains (Kosaki et al. 2002; Musante et al. 2003; Sarkozy et al. 2003; Zenker et al. 2004), suggesting that they are gain-of-function mutations with enhanced phosphatase activity. Mutations identified in leukemia and NS occurred at the same amino acid residues, but the type of substitutions rarely overlapped.
Recent studies have shown that the phosphatase activities of five mutations were elevated and that the phosphatase activities of two mutations (D61Y and E76K) identified in JMML cells were higher than that of the N308D mutation identified in NS (Tartaglia et al. 2003; Fragale et al. 2004). In this study, we performed phosphatase assays for 14 mutants identified in NS and leukemia to examine the presence of any functional differences between the mutations. Finally, we examined the functional consequences of the mutations in cultured cells.
Materials and methods
Patients
Forty-one patients with NS were recruited. The diagnosis of NS was evaluated by clinical geneticists based on the cardinal criteria delineated by Allanson (1987); characteristic faces including hypertelorism, down-slanting palpebral fissures and ptosis, webbed or short neck, chest deformity, cubitus vulgas, short stature, congenital heart defects, developmental delay, and cryptorchidism. Thirty-eight patients were sporadic cases, and three were familial cases. Bone marrow cells, peripheral blood, or cell lines were obtained from 29 leukemia cases without NS (seven ALL, 18 AML, one CML, two MDS, and one JMML). Eighteen AML patients were classified according to French-American-British classification (FAB); two with M0, one with M1, five with M2, three with M4, three with M5, one with M7, and three with unknown classification. This study was approved by the Ethics Committee of the Tohoku University School of Medicine.
Mutation analysis
After obtaining written informed consent, genomic DNA was isolated from the patients’ peripheral leukocytes, Epstein-Barr virus-transformed lymphoblastoid cells, or leukemia cells. Fifteen coding exons of PTPN11 from 41 NS patients and exons 3 and 13 from leukemia patients were sequenced. Each PTPN11 exon with flanking intronic sequences was amplified with primers based on GeneBank sequences (GeneBank accession no. NT 009775). The M13 reverse or forward sequence was added to the 5′ end of the polymerase chain reaction (PCR) primers for use as a sequencing primer. The PCR was performed in 30 μl of a solution containing 10 mM Tris–HCl (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, 0.2 mM dNTP, 10% (v/v) DMSO, 0.4 pmol of each primer, 100 ng genomic DNA, and 2.5 U of Taq DNA polymerase. The reaction condition consisted of 35 cycles of denaturation at 94°C for 15 s, annealing at 57°C for 15 s, and extension at 72°C for 30 s. The products were gel-purified and sequenced on an ABI PRISM 310 automated DNA sequencer (Applied Biosystems, Fostercity, CA, USA).
Construction of the plasmids
The human SHP-2 cDNA was PCR amplified by using the following primers: the 5′ primer included sequences encoding an EcoRI site followed by the FLAG M5 epitope (MDYKDDDDK) (Aoki et al. 2000). In addition, the 3′ primer had an EcoRI site. The amplified fragment was digested with EcoRI and subcloned into an EcoRI site of pBluescript KSII+ (Stratagene). Mutant constructs were generated using a Quickchange site-directed mutagenesis kit (Stratagene). All mutant and normal constructs were verified by sequencing. The full-length SHP-2 cDNA and mutants were digested with EcoRI and subcloned into an EcoRI site of the pCAGGS expression vector, a mammalian expression vector that has a cytomegalovirus enhancer, and a chicken β-actin promoter (Niwa et al. 1991).
Immune complex tyrosine phosphatase assay
The COS7 cells were purchased from the American Tissue Culture Collection (ATCC). The COS7 cells were maintained in DMEM containing 10% FCS, 100 U/ml penicillin, and 100 μg/ml streptomycin. Cells were seeded at 6×105 cells/6-cm dish, and 24 h later, 2.0 μg of pCAGGS vectors encoding one of the wild-type (WT) or mutant SHP-2 cDNAs were transfected using 8 μl of LipofectAMINE reagent and 12 μl of PLUS reagent (Invitrogen, Carlsbad, CA, USA). The E76A mutant identified somatically in leukemia cells (Tartaglia et al. 2003) was used as the positive control in each assay because this mutant was proved to have an increased phosphatase activity in a Xenopus experiment (O’Reilly et al. 2000). Cells were serum starved for 24 h, scraped, and collected by centrifugation after two washes with phosphate buffered saline (PBS). Lysates were prepared in 1 ml ice-cold RIPA buffer (50 mM Tris–HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1:100 protease inhibitor, and 1% Triton X) and incubated on ice for 15 min. The lysates were centrifuged at 14,000 g for 15 min at 4°C, and 1 mg of protein was used for immunoprecipitation. The FLAG-tagged WT SHP-2, and mutants were immunoprecipitated with anti-FLAG M5 antibody (Sigma, St Louis, Mosby, USA) for 2 h at 4°C. Immune complexes were collected by adding 50 μl of 50% protein G-Sepharose beads slurry (Amersham Biosciences Corp., Piscataway, NJ, USA) for 1 h at 4°C, washed twice with RIPA buffer, and then washed twice with phosphatase assay buffer (20 mM Hepes pH 7.5, 250 mM NaCl, 1 mM EDTA, 1 mM DTT).
Immune complex phosphatase assay was performed at 30°C for 30 min in 25 μl phosphatase assay buffer supplemented with 500 μM Src phosphopeptide (pTSTEPQpYQPGENL). The reaction was linear at the range of 0–1,000 μM Src. After brief centrifugation, supernatants were collected, added to 100 μl Malachite Green solution (Upstate Biotechnology), and incubated for 15 min at room temperature. The absorbance at 620 nm was measured, and the increase in the phosphatase activity was calculated by subtracting the background blank values (negative Src). Values for phosphate release were then obtained by comparing with the standard curve (linear at 0–2,000 pmol of phosphate). In order to measure the level of immunoprecipitated proteins, 50 μl 2× SDS sample buffer was added to the beads and boiled for 3 min. Twenty μl was used for SDS-PAGE followed by immunoblotting with anti-FLAG M5 antibody.
ERK (extracellular signal-regulated kinase) phosphorylation
Human embryonic kidney (HEK) 293 cells (ATCC) were maintained in DMEM containing 10% FCS, 100 U/ml penicillin, and 100 μg/ml of streptomycin. Cells were seeded at 6×105 cells/6-cm dish, and 24 h later, 3.6 μg of pCAGGS vectors encoding one of the WT or mutant SHP-2 cDNAs and 0.9 μg of pcDNA3 containing HA-tagged extracellular signal-regulated kinase (ERK)2 cDNA were cotransfected using 8 μl of LipofectAMINE reagent and 12 μl of PLUS reagent. Two major mutants in NS, D61N and E76D, were used for this study. Three hours later, the medium was changed to DMEM containing 10% FCS. At 48 h after transfection, cells were unstimulated or stimulated with 20 ng/ml of epidermal growth factor (EGF) for 5, 30, 60, 90, and 120 min.
Immunoprecipitation of HA-ERK2 was performed, as described previously (Takeda et al. 1999). The 293 cells were washed twice with PBS, and immediately frozen in liquid nitrogen. The cells were lysed on ice in 0.3 ml of ice-cold RIPA buffer [20 mM Tris–HCl (pH 7.5), 140 mM NaCl, 2.6 mM CaCl2, 1 mM MgCl2, 1% (v/v) NP-40 containing 1 mM phenylmethylsulfonyl fluoride (PMSF) and 3 μl phosphatase inhibitor (Sigma)]. The lysates were centrifuged at 14,000 g for 15 min at 4°C. For immunoprecipitation, the resultant supernatants were incubated with anti-HA antibody (clone12CA5, gifted by Dr. Matozaki) for 2 h at 4°C and then incubated with 30 μl of protein G-Sepharose beads (Amersham) for 1 h on a shaker. The beads were washed twice with 800 μl wash buffer [20 mM Tris–HCl (pH 7.5), 140 mM NaCl, 0.1% (v/v) Triton X-100] and boiled for 5 min in 2× SDS sample buffer. Immunoprecipitants were subjected to SDS-polyacrylamide gel electrophoresis (10% gel), transferred to nitrocellulose membrane, and probed with the indicated antisera. All the membranes were visualized using enhanced chemiluminescence (ECL) Western blotting Detection System Kit (Amersham Pharmacia Biotech, Chalfont, UK).
Reporter assay
One day before the transfection, the 293 cells were plated in 6-well plates with a density of 5×105 cells per well. Cells were transiently transfected using LipofectAMINE plus reagent with 1 μg pSREluc, 0.1 μg of pRLTKluc, and 1 μg of SHP-2 wild or mutant expression constructs. Four mutants in the N-SH2 domain from NS patients were introduced in 293 cells. Eighteen hours after transfection, the cells were serum-starved in DMEM for 24 h. For EGF stimulation, cells were unstimulated or stimulated with 20 ng/ml of EGF for 5 h. Cells were harvested in passive lysis buffer, and luciferase activity was assayed using a dual luciferase assay kit (Promega, Madison, WI, USA). Renilla luciferase expressed from pRLTKluc was used to normalize the transfection efficiency. The experiments were performed in triplicate. Data are shown as mean ± SEM. Statistical analysis was performed with the Statview 4.0 package (AVACUS Corporation, Berkeley, CA, USA).
Results
Mutation analysis in patients with NS
Sequencing analysis in 41 affected individuals with NS revealed nine different missense mutations in 16 patients (15 sporadic cases and one index case of familial origin) (Table 1). Ten patients had mutations in the N-SH2 domain (Y63C in three patients; D61N, Q79R, and T73I in two patients each; and F71I in one patient). The novel F71I mutation identified in patient 6 was inherited from his father, who exhibited only ptosis. The F71I mutation was not detected in 83 controls (data not shown), suggesting the mutation was not likely to be a polymorphism. Although the D61N, Y63C, and Q79R mutations were detected in more than two patients, their clinical findings, including facial anomalies and heart defects, varied significantly. Patient 8, who had the T73I mutation, exhibited JMML at 2 months of age. Her leukocytosis persisted for 20 years after remission. Four mutations in the PTP domains were detected in six patients. Interestingly, patient 15 with the S502T mutation had an episode of transient abnormal myelopoiesis at 3 months of age and was associated with neuroblastoma at 6 months, suggesting the extending characteristics of the mutation (Kondoh et al. 2003). The analysis of her parents’ DNA indicated the de novo mutation.
Phosphatase activities of mutants identified in NS
Structural analysis suggested that mutant proteins have an altered phosphatase activity. We transfected ten SHP-2 mutants, including nine mutants identified in NS patients in this study, and the E76A mutant as the positive control. Remarkable activation (six-fold to 12-fold compared with the activity of WT) was observed in the immune complexes derived from cells expressing the constructs carrying the D61N, F71I, or T73I mutation located in the N-SH2 domain (Fig.1). The activities of the D61N and F71I mutants were higher than that of the E76A mutant. Mild activation (two-fold to 3.9-fold) was observed in the immune complexes derived from cells transfected with constructs harboring the Y63C or Q79R mutation in the N-SH2 domain and constructs with the I282V, N308D, S502T, or M504V mutation in the PTP domain. Interestingly, all nine mutations had activated phosphatase activity.
Immune complex phosphatase assay of wild-type (WT) cDNA and mutants identified in Noonan syndrome (NS). The E76A mutant was used as a positive control in each assay. Results are expressed as mean ± SD obtained from three independent experiments. Asterisks indicate statistically significant differences (p < 0.01) in activity compared with that of WT. Fold activation was expressed as 1 for WT cDNA at each assay (upper panel). Immunoblot analysis of SHP-2 showed the comparative efficiency of immunoprecipitation (lower panel). HC heavy chain
PTPN11 mutation detection and phosphatase assay in childhood leukemia
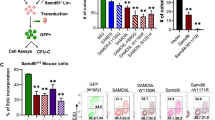
In order to screen PTPN11 mutation in leukemia cells, we sequenced exons 3 and 13 in the PTPN11 gene from genomic DNA extracted from 29 leukemic children. Two mutations were detected in three cases (Fig.2). The A72V mutation was identified in bone marrow from one case with AML (M0 according to FAB classification) and in bone marrow from one patient with MDS (refractory anemia with excess blasts) (Monosomy 7). The novel G503V (1508G>T) mutation was detected in a bone-marrow-derived cell line from a JMML patient. The phosphatase activity in an immune complex precipitated from cells overexpressing the A72V mutant showed a 5.2-fold increase and that from cells over expressing the G503V mutant showed a 1.4-fold increase compared with the WT cDNA (Fig.2b).
Phosphatase activity of mutants detected in childhood leukemia. a Localization of SHP-2 mutations analyzed in this study. Italic, mutations identified in this study. Parenthesis, number of patients identified. b Immune complex phosphatase assay using SHP-2 mutants identified in leukemia. Results are expressed as mean ± SD obtained from the three independent experiments. Asterisks indicate statistically significant differences in activity compared with that of wild type (WT) (* p < 0.01, ** p < 0.05). c, d Comparison of phosphatase activity in mutants derived from either NS or leukemia at amino acid positions 61 and 76. Results are expressed as mean ± SD obtained from the three independent experiments
In order to compare the phosphatase activity of mutations found in NS and leukemia at the same residue, we examined the phosphatase activity of two mutants occurring at D61 and E76. The activity of D61N (NS) was equivalent to that of D61Y (leukemia) (Fig.2c). In contrast, the activity of E76A (leukemia) was 4.1-fold that of E76D (NS) (Fig.2d).
a Effect of the D61N and E76D mutants on extracellular signal-regulated kinase (ERK)2 activation. Lysates were subjected to immunoprecipitation with anti-HA antibody (12CA5), and the immunoprecipitates were subjected to immunoblot analysis with anti-phospho-ERK antibody (upper panel). The aliquots of immunoprecipitates were used to confirm that equal amounts of HA-ERK2 were present in each lane (lower panel). Cell lysates from epidermal growth factor (EGF)-treated 293 cells were used as a positive control for phospho-ERK(1/2) immunoblotting (293). HC heavy chain. C459S catalytically inactive mutant. b EGF-stimulated transcription of the SRE reporter plasmid in cells expressing wild-type (WT), dominant negative SHP-2, and SHP-2 mutants in the N-SH2 domain. Each value represents luciferase activity in relative light units, which was normalized for Renilla luciferase activity. White bars, in the absence of EGF; filled bars, in the presence of EGF. Results are expressed as the mean ± SEM of the three independent experiments
Effects of mutants on MAPK pathway
Since SHP-2 has been known essentially as a positive regulator of the RAS/MAPK pathway (Feng 1999), we examined whether mutants activated ERK, a kind of MAPK, in the presence of EGF. In the absence and presence of EGF, the ERK activation was examined using an anti-phospho-ERK antibody that detected phosphorylated ERK. In cells expressing WT SHP-2 and ERK plasmids, the phosphorylation of ERK2 was maximum at 5 min after EGF treatment, and sustained activation persisted for 120 min (Fig.2a). In cells expressing the D61N or the E76D mutant, the maximum activation of ERK2 at 5 min was almost the same as in WT-transfected cells, but the activation decreased more rapidly. These results suggest that these two mutants do not promote the activation of the MAPK pathway in EGF-treated 293 cells.
In order to examine the downstream signaling pathway of ERK, we examined the activation of the serum-responsive element (SRE) within c-fos or other immediate early genes in EGF-treated 293 cells. As a key target of MAPK, ELK1 transcription factors phosphorylated by ERK bind to the SRE to initiate transcription (Bennett et al. 1996; Hakak and Martin 1999). Cells expressing the SRE-luciferase reporter gene and a catalytically inactive SHP-2 (C459S) showed a decreased induction of the luciferase gene in the presence of EGF (Fig.2b). Cotransfection of plasmids harboring mutants detected in NS had neither enhanced SRE activity in the unstimulated state nor enhanced fold induction in the presence of EGF. These results suggest that the SHP-2 mutants did not promote the MAPK pathway in the EGF-treated 293 cells whereas they showed enhanced phosphatase activity in vitro.
Discussion
In this study, we identified nine mutations, including a novel F71I mutation, in 16 of 41 Japanese patients with NS and two mutations, including a novel G503V mutation, in three of 29 patients with childhood leukemia. Immune complex phosphatase assays showed that the phosphatase activity in ten SHP-2 mutants detected in NS and four SHP-2 mutants detected in leukemia was elevated to various degrees. Mutations in the PTPN11 gene were detected in 39% of the patients clinically diagnosed with NS. This result is comparable to that of previous studies (Tartaglia et al. 2002; Musante et al. 2003). Previous genotype–phenotype correlation studies have shown that pulmonary stenosis was associated with PTPN11 mutations whereas cardiomyopathy was more common in patients without PTPN11 mutation (Tartaglia et al. 2002; Zenker et al. 2004). In our study, however, no significant difference was observed (data not shown). Although other abnormalities than exonic mutations in PTPN11 have not been excluded, further studies to identify the gene(s) associated with NS will pose a formidable challenge in establishing the clinical entity of NS.
The role of PTPN11 mutations in the pathogenesis of hematologic malignancy remains to be elucidated. In the previous and the present studies, 87% of the mutations found in leukemia cells were identified in the N-SH2 domain, and these were primarily restricted to residues D61, A72, and E76 (Tartaglia et al. 2003; Loh et al. 2004; Tartaglia et al. 2004). To examine whether the degrees of phosphatase activity in mutations at specific residues are associated with leukemia, we compared the phosphatase activity at each affected residue examined in this study, as well as the published phenotype (Table 2a). Variation of each assay was normalized as relative phosphatase activity; activity of WT cDNA was taken as one-fold, and that of E76A was taken as ten-fold. Mutations at codons 61, 71, 72, and 76, in which highest activity at each codon was ten or more, were frequently identified in patients with leukemia, including those with JMML, MDS, AML, and ALL (Table 2a). In NS patients, activities of the most common mutations at N308 and Y63 were two-fold to three-fold. The phosphatase activity of the T73I mutation, which is frequently detected in NS patients associated with JMML, showed an intermediate value of 6.0. Mutations at residues in which activities were below six were rarely detected in leukemia cells. These results suggest that mutations at codons 61, 71, 72, and 76, in which highest activity at each codon was ten or more, were associated with leukemia (Table 2b; χ2=97.0, p < 0.001).
Tartaglia et al. (2003) recently showed that the phosphatase activities of two mutants (D61Y and E76K) identified in JMML patients were higher than that of the N308D mutation identified in NS. They put forth a hypothesis explaining the manner in which the two different phenotypes were determined: somatic mutations identified in leukemia have stronger molecular defects and could cause embryonic lethality if they occur in the germline. In contrast, the defects in mutations found in NS patients were milder and insufficient for the development of leukemia. In our experiments, the relative phosphatase activities of D61N (12.1) and F71I (11.8) associated with NS were higher than those of E76A (10) and G503V (2.7) found in leukemia. A comparison of the different substitutions at the same residues showed that the phosphatase activity of D61N (NS) was equivalent to that of D61Y (leukemia). Although Src might not be the physiological substrate of SHP-2 at the developmental stage and leukemogenesis, our results suggest that high phosphatase activity in mutants may not necessarily be associated with embryonic lethality. Further biological studies will be required to explore the mechanisms involved in the development of NS or leukemia.
The F71I mutation in a typical NS patient was inherited from the father, who exhibited only ptosis. The F71I mutation was not detected in 83 controls, and a further functional analysis showed an elevated phosphatase activity. These observations support the notion that the F71I mutation plays a pathogenic role. One possible explanation for the discrepancy between the clinical and in vitro data is the phenotypic change with age. Allanson et al. (1958) documented the marked change in phenotype with age and reported that manifestations in adults tend to be subtle. Although heart defects and other medical problems had not been observed in the father, it is possible that his facial abnormalities during childhood had been more prominent. Although SHP-2 was essentially expressed in adult organs (Ahmad et al. 1993), in this case, the SHP-2 mutant could have affected specific signaling pathways at the developmental stage. Another possible explanation for his subtle phenotype was due to the poor penetrance of the mutation. Since the genotype–phenotype correlation was not observed in a previous report (Zenker et al. 2004), it is possible that genetic factors other than PTPN11 are associated with the phenotype of NS.
A recent paper showed that three PTPN11 mutations were identified in 89 primary neuroblastomas (Bentires-Alj et al. 2004). Two mutations (E69K and T507K) were identified as somatic mutations. Y62C was identified in both tumor and normal adrenal tissues, suggesting that the patient might be NS. In our study, the patient with S502T developed leukocytosis at 3 months of age and neuroblastoma at 6 months of age (Kondoh et al. 2003). Previous reports showed that the insulin-like growth factor or the hepatocyte growth factor stimulated the proliferation or invasion of neuroblastoma cells (Zumkeller and Schwab 1999; Hecht et al. 2004). It is known that SHP-2 regulates these growth-factor signal pathways (Neel et al. 2003). Although the exact mechanisms remain unknown, it is possible that the activated mutations were associated with the pathogenesis of neuroblastoma. It is important to note that the neuroblastoma in the NS patient with the S502T mutation spontaneously disappeared (Kondoh et al. 2003). A similar phenomenon was observed in NS patients associated with JMML, most of which spontaneously resolved (Choong et al. 1999).
A previous report showed that SHP-2 positively regulated the EGF-mediated ERK activation, ELK-1 transactivation, and cell cycle progression in 293 cells (Bennett et al. 1996). In our experiments, two mutations associated with NS failed to promote the RAS/MAPK pathway in the EGF-treated 293 cells. It is controversial as to whether constitutive active mutants lead to enhanced RAS/MAPK pathway. In Xenopus, it was shown that the D61A and E76A mutants induced an elongation of the animal cap. However, this elongation was accompanied by minimal activation of MAPK (O’Reilly et al. 2000). Fragale et al. (2004) reported that mutations such as A72V, I282V, and N308D detected in the NS patients caused prolonged binding of GAB1, a binding partner of SHP-2, and sustained ERK activation when GAB1 was cointroduced in EGF-treated COS cells. In contrast, ERK was not hyperactivated in Ba/F3 cells expressing mutants associated with JMML (D61G and E76K) although these cells showed enhanced growth-factor-independent survival. A recent study analyzing knock-in mice expressing the D61G mutation showed cell- and pathway-specific EKR activation in the developing mice (Araki et al. 2004). These experiments and our data suggest that the manner in which the constitutive active mutants regulate their downstream signaling pathway(s) depends on cell types and signaling pathways.
In conclusion, we showed that the phosphatase activity of 14 mutations identified in NS and leukemia was elevated to various degrees. High phosphatase activity in mutations at codons 61, 71, 72, and 76 was associated with leukemogenesis. A further analysis of the downstream signals of the mutants would provide the key to elucidate the role of PTPN11 mutations in patients with NS and leukemia.
References
Ahmad S, Banville D, Zhao Z, Fischer EH, Shen SH (1993) A widely expressed human protein-tyrosine phosphatase containing src homology 2 domains. Proc Natl Acad Sci USA 90:2197–2201
Allanson JE (1987) Noonan syndrome. J Med Genet 24:9–13
Allanson JE, Hall JG, Hughes HE, Preus M, Witt RD (1985) Noonan syndrome: the changing phenotype. Am J Med Genet 21:507–514
Aoki Y, Huang Z, Thomas SS, Bhide PG, Huang I, Moskowitz MA, Reeves SA (2000) Increased susceptibility to ischemia-induced brain damage in transgenic mice overexpressing a dominant negative form of SHP2. Faseb J 14:1965–1973
Araki T, Mohi MG, Ismat FA, Bronson RT, Williams IR, Kutok JL, Yang W, Pao LI, Gilliland DG, Epstein JA, Neel BG (2004) Mouse model of Noonan syndrome reveals cell type- and gene dosage-dependent effects of Ptpn11 mutation. Nat Med 10:849–857
Bennett AM, Hausdorff SF, O’Reilly AM, Freeman RM, Neel BG (1996) Multiple requirements for SHPTP2 in epidermal growth factor-mediated cell cycle progression. Mol Cell Biol 16:1189–1202
Bentires-Alj M, Paez JG, David FS, Keilhack H, Halmos B, Naoki K, Maris JM, Richardson A, Bardelli A, Sugarbaker DJ, Richards WG, Du J, Girard L, Minna JD, Loh ML, Fisher DE, Velculescu VE, Vogelstein B, Meyerson M, Sellers WR, Neel BG (2004) Activating mutations of the noonan syndrome-associated SHP2/PTPN11 gene in human solid tumors and adult acute myelogenous leukemia. Cancer Res 64:8816–8820
Choong K, Freedman MH, Chitayat D, Kelly EN, Taylor G, Zipursky A (1999) Juvenile myelomonocytic leukemia and Noonan syndrome. J Pediatr Hematol Oncol 21:523–527
Feng GS (1999) Shp-2 tyrosine phosphatase: signaling one cell or many. Exp Cell Res 253:47–54
Fragale A, Tartaglia M, Wu J, Gelb BD (2004) Noonan syndrome-associated SHP2/PTPN11 mutants cause EGF-dependent prolonged GAB1 binding and sustained ERK2/MAPK1 activation. Hum Mutat 23:267–277
Hakak Y, Martin GS (1999) Cas mediates transcriptional activation of the serum response element by Src. Mol Cell Biol 19:6953–6962
Hecht M, Papoutsi M, Tran HD, Wilting J, Schweigerer L (2004) Hepatocyte growth factor/c-Met signaling promotes the progression of experimental human neuroblastomas. Cancer Res 64:6109–6118
Hof P, Pluskey S, Dhe-Paganon S, Eck MJ, Shoelson SE (1998) Crystal structure of the tyrosine phosphatase SHP-2. Cell 92:441–450
Kondoh T, Ishii E, Aoki Y, Shimizu T, Zaitsu M, Matsubara Y, Moriuchi H (2003) Noonan syndrome with leukaemoid reaction and overproduction of catecholamines: a case report. Eur J Pediatr 162:548–549
Kosaki K, Suzuki T, Muroya K, Hasegawa T, Sato S, Matsuo N, Kosaki R, Nagai T, Hasegawa Y, Ogata T (2002) PTPN11 (protein-tyrosine phosphatase, nonreceptor-type 11) mutations in seven Japanese patients with Noonan syndrome. J Clin Endocrinol Metab 87:3529–3533
Loh ML, Vattikuti S, Schubbert S, Reynolds MG, Carlson E, Lieuw KH, Cheng JW, Lee CM, Stokoe D, Bonifas JM, Curtiss NP, Gotlib J, Meshinchi S, Le Beau MM, Emanuel PD, Shannon KM (2004) Mutations in PTPN11 implicate the SHP-2 phosphatase in leukemogenesis. Blood 103:2325–2331
Maheshwari M, Belmont J, Fernbach S, Ho T, Molinari L, Yakub I, Yu F, Combes A, Towbin J, Craigen WJ, Gibbs R (2002) PTPN11 mutations in Noonan syndrome type I: detection of recurrent mutations in exons 3 and 13. Hum Mutat 20:298–304
Mendez HM, Opitz JM (1985) Noonan syndrome: a review. Am J Med Genet 21:493–506
Musante L, Kehl HG, Majewski F, Meinecke P, Schweiger S, Gillessen-Kaesbach G, Wieczorek D, Hinkel GK, Tinschert S, Hoeltzenbein M, Ropers HH, Kalscheuer VM (2003) Spectrum of mutations in PTPN11 and genotype-phenotype correlation in 96 patients with Noonan syndrome and five patients with cardio-facio-cutaneous syndrome. Eur J Hum Genet 11:201–206
Neel BG, Gu H, Pao L (2003) The ‘Shp’ing news: SH2 domain-containing tyrosine phosphatases in cell signaling. Trends Biochem Sci 28:284–293
Niwa H, Yamamura K, Miyazaki J (1991) Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 108:193–199
O’Reilly AM, Pluskey S, Shoelson SE, Neel BG (2000) Activated mutants of SHP-2 preferentially induce elongation of Xenopus animal caps. Mol Cell Biol 20:299–311
Sarkozy A, Conti E, Seripa D, Digilio MC, Grifone N, Tandoi C, Fazio VM, Di Ciommo V, Marino B, Pizzuti A, Dallapiccola B (2003) Correlation between PTPN11 gene mutations and congenital heart defects in Noonan and LEOPARD syndromes. J Med Genet 40:704–708
Schollen E, Matthijs G, Gewillig M, Fryns JP, Legius E (2003) PTPN11 mutation in a large family with Noonan syndrome and dizygous twinning. Eur J Hum Genet 11:85–88
Servidei T, Aoki Y, Lewis SE, Symes A, Fink JS, Reeves SA (1998) Coordinate regulation of STAT signaling and c-fos expression by the tyrosine phosphatase SHP-2. J Biol Chem 273:6233–6241
Takeda H, Matozaki T, Takada T, Noguchi T, Yamao T, Tsuda M, Ochi F, Fukunaga K, Inagaki K, Kasuga M (1999) PI 3-kinase gamma and protein kinase C-zeta mediate RAS-independent activation of MAP kinase by a Gi protein-coupled receptor. EMBO J 18:386–395
Tartaglia M, Mehler EL, Goldberg R, Zampino G, Brunner HG, Kremer H, van der Burgt I, Crosby AH, Ion A, Jeffery S, Kalidas K, Patton MA, Kucherlapati RS, Gelb BD (2001) Mutations in PTPN11, encoding the protein tyrosine phosphatase SHP-2, cause Noonan syndrome. Nat Genet 29:465–468
Tartaglia M, Kalidas K, Shaw A, Song X, Musat DL, van der Burgt I, Brunner HG, Bertola DR, Crosby A, Ion A, Kucherlapati RS, Jeffery S, Patton MA, Gelb BD (2002) PTPN11 mutations in Noonan syndrome: molecular spectrum, genotype-phenotype correlation, and phenotypic heterogeneity. Am J Hum Genet 70:1555–1563
Tartaglia M, Niemeyer CM, Fragale A, Song X, Buechner J, Jung A, Hahlen K, Hasle H, Licht JD, Gelb BD (2003) Somatic mutations in PTPN11 in juvenile myelomonocytic leukemia, myelodysplastic syndromes and acute myeloid leukemia. Nat Genet 34:148–150
Tartaglia M, Martinelli S, Cazzaniga G, Cordeddu V, Iavarone I, Spinelli M, Palmi C, Carta C, Pession A, Arico M, Masera G, Basso G, Sorcini M, Gelb BD, Biondi A (2004) Genetic evidence for lineage-related and differentiation stage-related contribution of somatic PTPN11 mutations to leukemogenesis in childhood acute leukemia. Blood 104:307–313
Yoshida R, Hasegawa T, Hasegawa Y, Nagai T, Kinoshita E, Tanaka Y, Kanegane H, Ohyama K, Onishi T, Hanew K, Okuyama T, Horikawa R, Tanaka T, Ogata T (2004) Protein-tyrosine phosphatase, nonreceptor type 11 mutation analysis and clinical assessment in 45 patients with noonan syndrome. J Clin Endocrinol Metab 89:3359–3364
Zenker M, Buheitel G, Rauch R, Koenig R, Bosse K, Kress W, Tietze HU, Doerr HG, Hofbeck M, Singer H, Reis A, Rauch A (2004) Genotype-phenotype correlations in Noonan syndrome. J Pediatr 144:368–374
Zumkeller W, Schwab M (1999) Insulin-like growth factor system in neuroblastoma tumorigenesis and apoptosis: potential diagnostic and therapeutic perspectives. Horm Metab Res 31:138–141
Acknowledgements
The authors thank the patients and their families who participated in this study and doctors who referred the cases. We are grateful to Dr. Takashi Matozaki, Gunma University, for supplying pcDNA-HAERK2 plasmid and anti-HA antibody, and Dr. Jun-ichi Miyazaki, Osaka University, for supplying the pCAGGS expression vector. We thank Kumi Kato, Dr. Xue Yang, and Dr. Yoshio Takahashi for their technical assistance. This work was supported by Grants-in-Aid from the Ministry of Education, Culture, Sports, Science and Technology of Japan (No.15790172) and the Uehara Memorial Foundation Grant-in-Aid for Scientific Research to YA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Niihori, T., Aoki, Y., Ohashi, H. et al. Functional analysis of PTPN11/SHP-2 mutants identified in Noonan syndrome and childhood leukemia. J Hum Genet 50, 192–202 (2005). https://doi.org/10.1007/s10038-005-0239-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-005-0239-7
Keywords
This article is cited by
-
Ganglioglioma with adverse clinical outcome and atypical histopathological features were defined by alterations in PTPN11/KRAS/NF1 and other RAS-/MAP-Kinase pathway genes
Acta Neuropathologica (2023)
-
Molecular and environmental characterization of Noonan syndrome in Morocco reveals a significant association with consanguinity and advanced parental age
Egyptian Journal of Medical Human Genetics (2020)
-
A PTPN11 mutation in a woman with Noonan syndrome and protein-losing enteropathy
BMC Gastroenterology (2020)
-
RAS nucleotide cycling underlies the SHP2 phosphatase dependence of mutant BRAF-, NF1- and RAS-driven cancers
Nature Cell Biology (2018)
-
Structural insights into Noonan/LEOPARD syndrome-related mutants of protein-tyrosine phosphatase SHP2 (PTPN11)
BMC Structural Biology (2014)