Abstract
Introduction
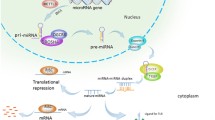
MicroRNAs (miRNAs) are small (∽22 nucleotides) regulatory RNAs which play fundamental roles in many human diseases, including cancer. There is no report on the miRNA expression profile of retinoblastoma.
Methods
This work was undertaken to identify differentially expressed miRNAs in human retinoblastoma tissues by microRNA microarray technique, and some miRNAs were verified using northern blot analysis and the in situ hybridization method.
Results
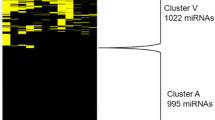
A cluster of microRNAs was identified as highly expressed in retinoblastoma, including hsa-miR-494, hsa-let-7e, hsa-miR-513–1, hsa-miR-513–2, hsa-miR-518c*, hsa-miR-129–1, hsa-miR-129–2, hsa-miR-198, hsa-miR-492, hsa-miR-498, hsa-miR-320, hsa-miR-503, and hsa-miR-373*.
Conclusion
These miRNAs are the first to be reported for human retinoblastoma and may play significant roles in regulating tumor genesis.



Similar content being viewed by others
References
Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE, Zhai Y, Giordano TJ, Qin ZS, Moore BB, MacDougald OA, Cho KR, Fearon ER (2007) p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol 17:1298–1307
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M, Croce CM (2004) Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 101:2999–3004
Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE, Iorio MV, Visone R, Sever NI, Fabbri M, Iuliano R, Palumbo T, Pichiorri F, Roldo C, Garzon R, Sevignani C, Rassenti L, Alder H, Volinia S, Liu CG, Kipps TJ, Negrini M, Croce CM (2005) A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 353:1793–1801
Cavenee WK, Hansen MF, Nordenskjold M, Kock E, Maumenee I, Squire JA, Phillips RA, Gallie BL (1985) Genetic origin of mutations predisposing to retinoblastoma. Science 228:501–503
Chakraborty S, Khare S, Dorairaj SK, Prabhakaran VC, Prakash DR, Kumar A (2007) Identification of genes associated with tumorigenesis of retinoblastoma by microarray analysis. Genomics 90:344–353
Chan JA, Krichevsky AM, Kosik KS (2005) MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 65:6029–6033
Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH, Feldmann G, Yamakuchi M, Ferlito M, Lowenstein CJ, Arking DE, Beer MA, Maitra A, Mendell JT (2007) Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 26:745–752
Chen D, Gallie BL, Squire JA (2001) Minimal regions of chromosomal imbalance in retinoblastoma detected by comparative genomic hybridization. Cancer Genet Cytogenet 129:57–63
Chen D, Pajovic S, Duckett A, Brown VD, Squire JA, Gallie BL (2002) Genomic amplification in retinoblastoma narrowed to 0.6 megabase on chromosome 6p containing a kinesin-like gene, RBKIN. Cancer Res 62:967–971
Chen D, Livne-bar I, Vanderluit JL, Slack RS, Agochiya M, Bremner R (2004) Cell-specific effects of RB or RB/p107 loss on retinal development implicate an intrinsically death-resistant cell-of-origin in retinoblastoma. Cancer Cell 5:539–551
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M, Wojcik SE, Aqeilan RI, Zupo S, Dono M, Rassenti L, Alder H, Volinia S, Liu CG, Kipps TJ, Negrini M, Croce CM (2005) miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 102:13944–13949
Corney DC, Flesken-Nikitin A, Godwin AK, Wang W, Nikitin AY (2007) MicroRNA-34b and MicroRNA-34c are targets of p53 and cooperate in control of cell proliferation and adhesion-independent growth. Cancer Res 67:8433–8438
Friend SH, Bernards R, Rogelj S, Weinberg RA, Rapaport JM, Albert DM, Dryja TP (1986) A human DNA segment with properties of the gene that predisposes to retinoblastoma and osteosarcoma. Nature 323:643–646
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, Powers S, Cordon-Cardo C, Lowe SW, Hannon GJ, Hammond SM (2005) A microRNA polycistron as a potential human oncogene. Nature 435:828–833
Huppi K, Volfovsky N, Mackiewicz M, Runfola T, Jones TL, Martin SE, Stephens R, Caplen NJ (2007) MicroRNAs and genomic instability. Semin Cancer Biol 17:65–73
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S, Magri E, Pedriali M, Fabbri M, Campiglio M, Menard S, Palazzo JP, Rosenberg A, Musiani P, Volinia S, Nenci I, Calin GA, Querzoli P, Negrini M, Croce CM (2005) MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65:7065–7070
Karali M, Peluso I, Marigo V, Banfi S (2007) Identification and characterization of microRNAs expressed in the mouse eye. Invest Ophthalmol Vis Sci 48:509–515
Kluiver J, Poppema S, de Jong D, Blokzijl T, Harms G, Jacobs S, Kroesen BJ, van den Berg A (2005) BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas. J Pathol 207:243–249
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, Johnson JM (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433:769–773
Meng F, Henson R, Wehbe-Janek H, Smith H, Ueno Y, Patel T (2007) The MicroRNA let-7a modulates interleukin-6-dependent STAT-3 survival signaling in malignant human cholangiocytes. J Biol Chem 282:8256–8264
Michael MZ, SM OC, van Holst Pellekaan NG, Young GP, James RJ (2003) Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 1:882–891
Ryan DG, Oliveira-Fernandes M, Lavker RM (2006) MicroRNAs of the mammalian eye display distinct and overlapping tissue specificity. Mol Vis 12:1175–1184
Sevignani C, Calin GA, Nnadi SC, Shimizu M, Davuluri RV, Hyslop T, Demant P, Croce CM, Siracusa LD (2007) MicroRNA genes are frequently located near mouse cancer susceptibility loci. Proc Natl Acad Sci USA 104:8017–8022
Strong LC (1977) Genetic etiology of cancer. Cancer 40:438–444
Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H, Harano T, Yatabe Y, Nagino M, Nimura Y, Mitsudomi T, Takahashi T (2004) Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 64:3753–3756
Tam W, Dahlberg JE (2006) miR-155/BIC as an oncogenic microRNA. Genes Chromosomes Cancer 45:211–212
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F, Visone R, Iorio M, Roldo C, Ferracin M, Prueitt RL, Yanaihara N, Lanza G, Scarpa A, Vecchione A, Negrini M, Harris CC, Croce CM (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103:2257–2261
Wightman B, Ha I, Ruvkun G (1993) Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75:855–862
Yang H, Kong W, He L, Zhao JJ, O’Donnell JD, Wang J, Wenham RM, Coppola D, Kruk PA, Nicosia SV, Cheng JQ (2008) MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res 68:425–433
Zhang B, Pan X, Cobb GP, Anderson TA (2007) microRNAs as oncogenes and tumor suppressors. Dev Biol 302:1–12
Zhao JJ, Hua YJ, Sun DG, Meng XX, Xiao HS, Ma X (2006) Genome-wide microRNA profiling in human fetal nervous tissues by oligonucleotide microarray. Childs Nerv Syst 22:1419–1425
Zhao JJ, Sun DG, Wang J, Liu SR, Zhang CY, Zhu MX, Ma X (2008) Retinoic acid downregulates microRNAs to induce abnormal development of spinal cord in spina bifida rat model. Childs Nerv Syst 24:485–492
Acknowledgements
We thank the patients, the normal individuals, and their family members for taking part in this study. This work was supported in part by the National Basic Research Program of China (2007CB511905), the National Infrastructure Program of Chinese Genetic Resources (2006DKA21301), National Natural science Foundation of China (30872843), the Program for Innovative Researcher Team in Science and Technology in Fujian Province University (FMU-RT001) and Fujian Provincial Natural Science Foundation (C0520003, C0810018).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Jian-Jun Zhao and Juhua Yang contribute equally to the article.
Rights and permissions
About this article
Cite this article
Zhao, JJ., Yang, J., Lin, J. et al. Identification of miRNAs associated with tumorigenesis of retinoblastoma by miRNA microarray analysis. Childs Nerv Syst 25, 13–20 (2009). https://doi.org/10.1007/s00381-008-0701-x
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00381-008-0701-x